MIRTK Release 2.0
The next major release of MIRTK is now available on GitHub!
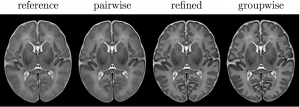
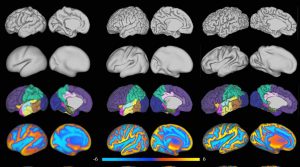
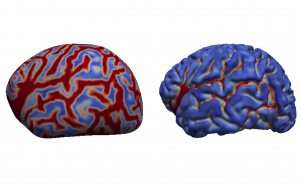
This release version was used in the experiments of my PhD thesis, and the “Unbiased construction of a temporally consistent morphological atlas of neonatal brain development” (doi:10.1101/251512). It also includes the tools used in the minimal processing pipeline of the dHCP (doi:10.1101/125526).